Description:
Enables high-yield production of unstable, post-translationally modified proteins by protecting them during expression and gentle release
Background
Intrinsically disordered proteins (IDPs), particularly those carrying labile post-translational modifications (PTMs), are difficult to produce recombinantly due to degradation, aggregation, and loss of modification during expression. Existing strategies often yield low quantities, heterogeneous products, or require harsh chemical treatments that damage sensitive PTMs, limiting their use in research, diagnostics, and drug discovery.
Technology Description
This technology (PIInBody) is a modular fusion-tag expression platform that directs proteins of interest into inclusion bodies during recombinant expression, protecting them from degradation and enzymatic loss of post-translational modifications. Unlike conventional inclusion body tags, the system is engineered to allow rapid refolding of the fusion tag and highly specific protease-mediated release of the target protein under mild, native conditions in less than 1 hr.
The platform is compatible with site-specific incorporation of authentic or non-hydrolyzable PTMs (e.g., phosphoserine) using genetic code expansion, with isotopic labeling, enabling production of homogeneous, precisely modified protein proteoforms for a variety of applications. The technology has been demonstrated at milligram scale with highly unstable intrinsically disordered proteins, including full-length tau and BAD, while preserving PTM integrity.
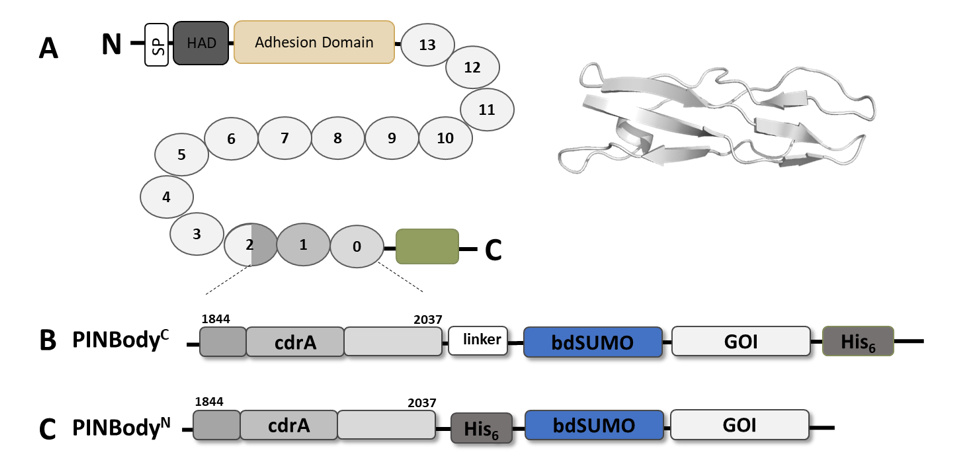
Design of PINBody fusion tags using the CdrA adhesion protein. (a) Domain organization of the full-length CdrA protein from Pseudomonas aeruginosa, including an N-terminal signal peptide (SP), hemagglutination activity domain (HAD), adhesion domain, and 14 extracellular β-strand-rich repeats, followed by a C-terminal periplasmic tail (green). Structure of repeat 0 was predicted by AlphaFold3 and visualized in PyMOL (right). Residues 1844–2037 comprising repeat 0, repeat 1, and part of repeat 2 were selected for the PINBody tag. (b) Schematic of the PINBodyC architecture, in which CdrA (residues 1844–2037) is fused to a flexible linker, bdSUMO, a gene of interest (GOI), and a C-terminal His₆ tag. (c) In the PINBodyN variant, the His₆ tag is placed between CdrA and bdSUMO, enabling purification and subsequent bdSENP1 cleavage to yield tag-free GOIs. Amino acid sequences for PINBodyC and PINBodyN are provided in Supplement information.
Features & Benefits
- Preserves labile post-translational modifications during expression
- Enables high-yield production of otherwise unstable proteins
- Avoids harsh chemical cleavage and denaturing conditions
- Produces homogeneous, site-specifically modified proteins
- Scalable and compatible with standard bacterial expression systems, including isotopic (e.g, 15N) labeling
- Flexible architecture supports multiple purification strategies
Applications
- Structural biology (e.g., NMR, mass spectrometry)
- Neurodegeneration and cancer research
- Drug discovery and screening assays
- Diagnostic standards and assay development
- Antibody and binder development against PTM-specific epitopes
- Research reagent production
Status
US Patent Application No. 63/975,085